Problem with Ramachandran Plot
-
After loading a PDB file (1CRN for example) and selecting Ramachandran Plot (Menu Biology and More), i can't see any residues on the map ! Selecting a single residue or the whole protein is not changing anything.
Any idea ? -
Dear @Emmanuel ,
And what happens if you click the Update button placed below the Ramachandran plot? For me, it works with PDB 1CRN (see the image below).
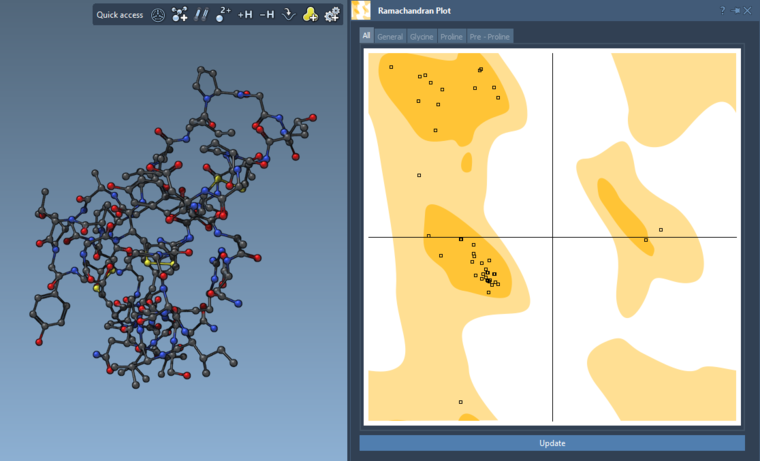
-
Thanks. But if 1CRN is already loaded, i expect to see the ramachandran plot when i open it the first time ! This is not the case. I have to click update even the first time to see it.
If i select a residue and click update, i see all protein residues not only the one selected. Is it correct ? -
If you select a residue or a set of residues and then click Update in the Ramachandran plot then it will have only these residues depicted.
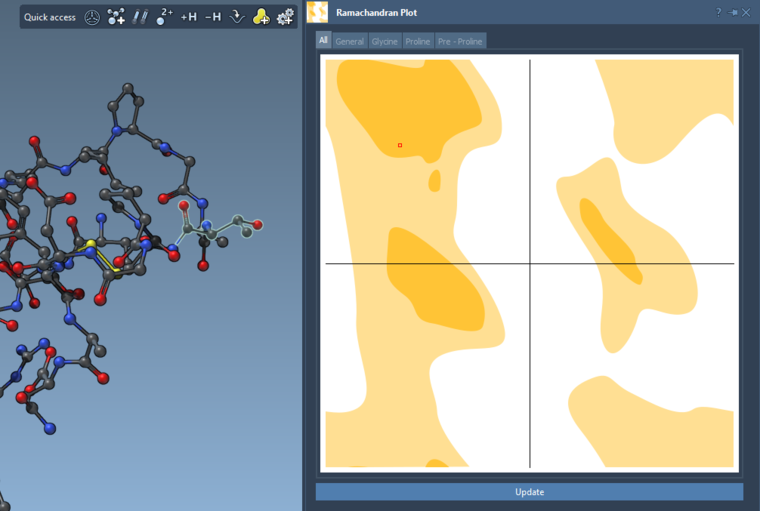
If you show for the whole structure, then you can also see the selected residues on the plot highlighted in red.
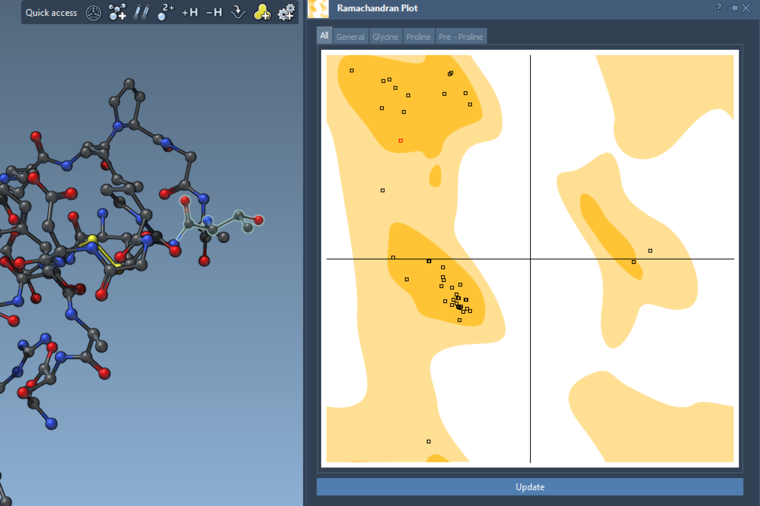
And you can interactively modify the backbone angles via this plot:
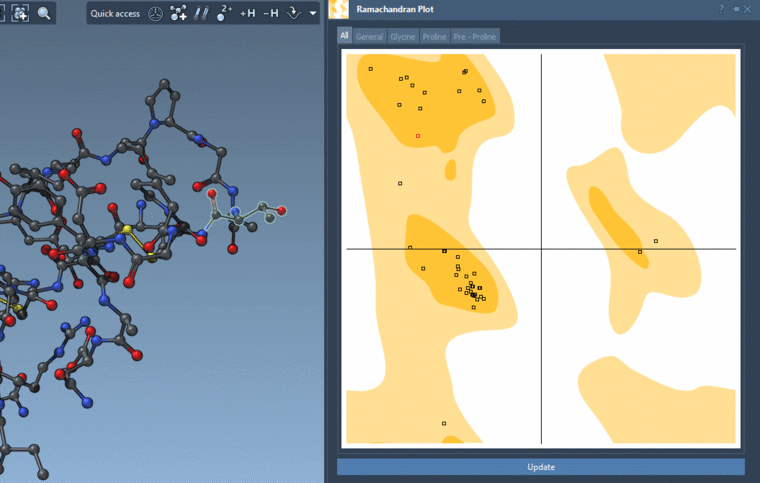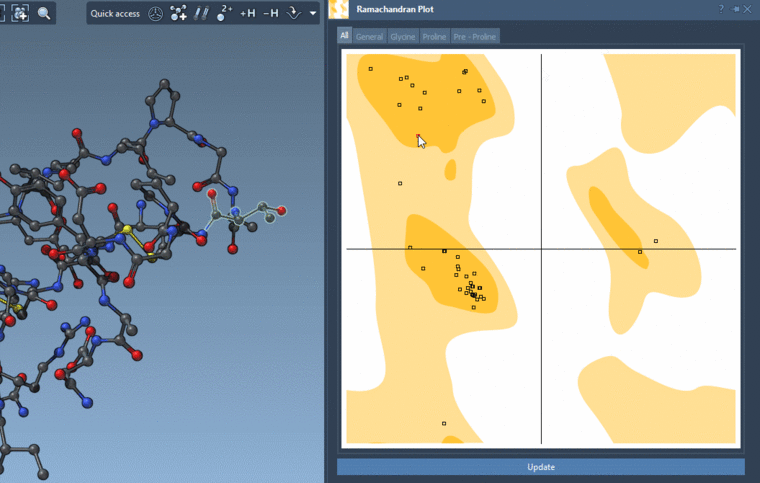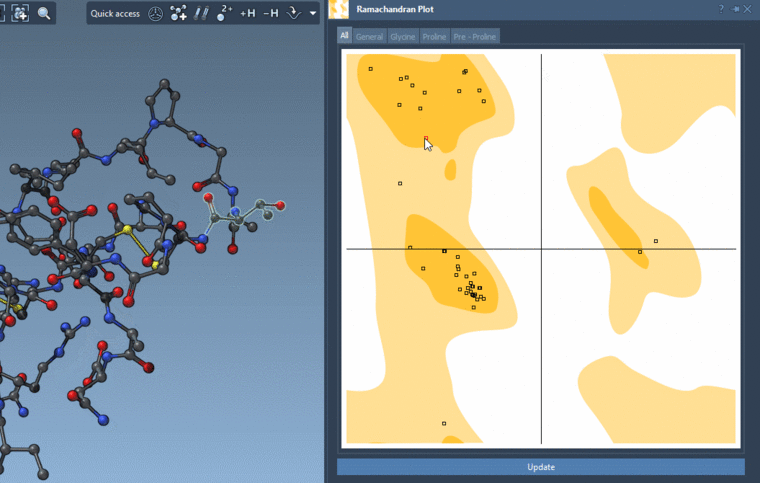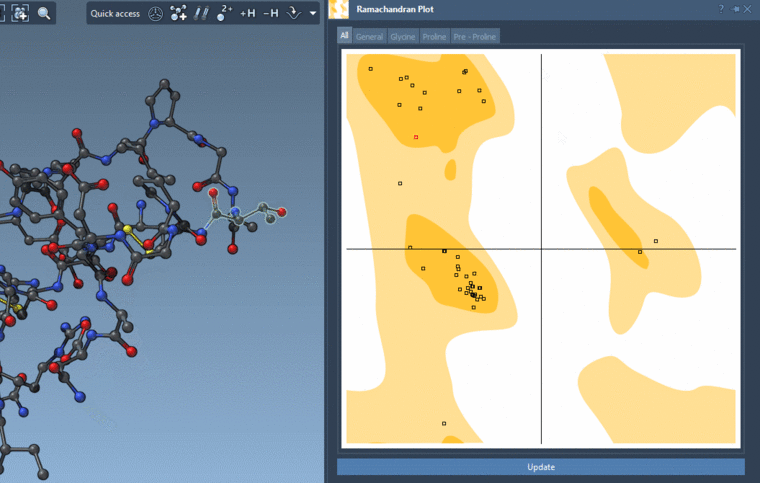
-
No. It's not working.
If i select a residue or a set of residues and then click Update in the Ramachandran plot i see ALL residues. -
Could you please describe how you perform the selection? There are two main possibilities apart from performing selections using the Node Specification Language:
-1- Selecting residues using the Document view.
-2- Selecting residues in the Viewport using the Select editor with the Selection filter set to Residues. If you are selecting in the Viewport with the Selection filter set to something e.g. Atoms and bonds then it will be atoms and bonds which are selected and not the residues.Then you can click Update in the Ramachandran plot and it will show data only for the selected residues if there are residues which are selected else it will show data for all the residues.
See the gif below for the both ways of selecting.
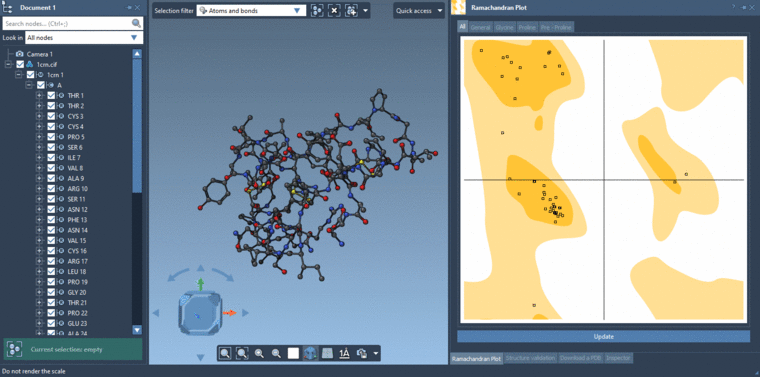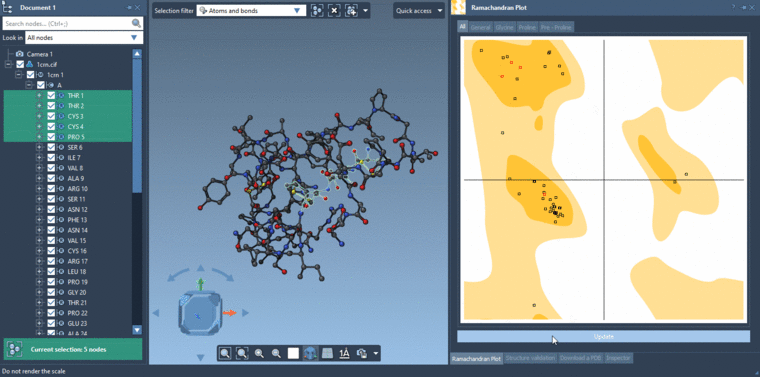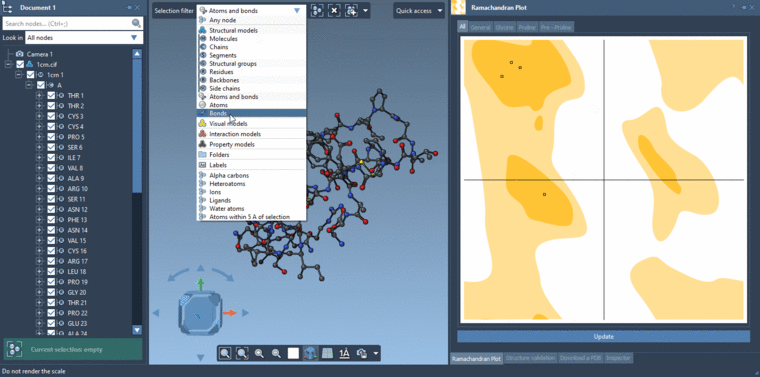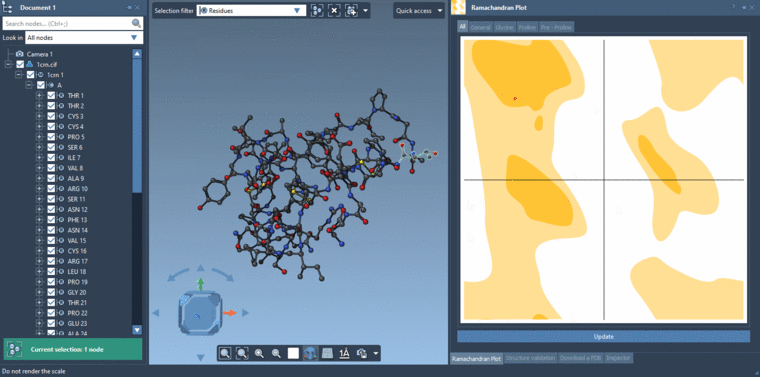
-
OK. That's the point. It's working if the Selection filter is set to Residues.
It is confusing. If i select Ramachandran Plot, is it possible to switch automatically to Residues instead of Atoms and bond by default ? -
Currently, it is not possible because the selection filter functionality is not exposed to external elements. We will update it in the upcoming release of SAMSON.