Getting atoms inside a .stl object
-
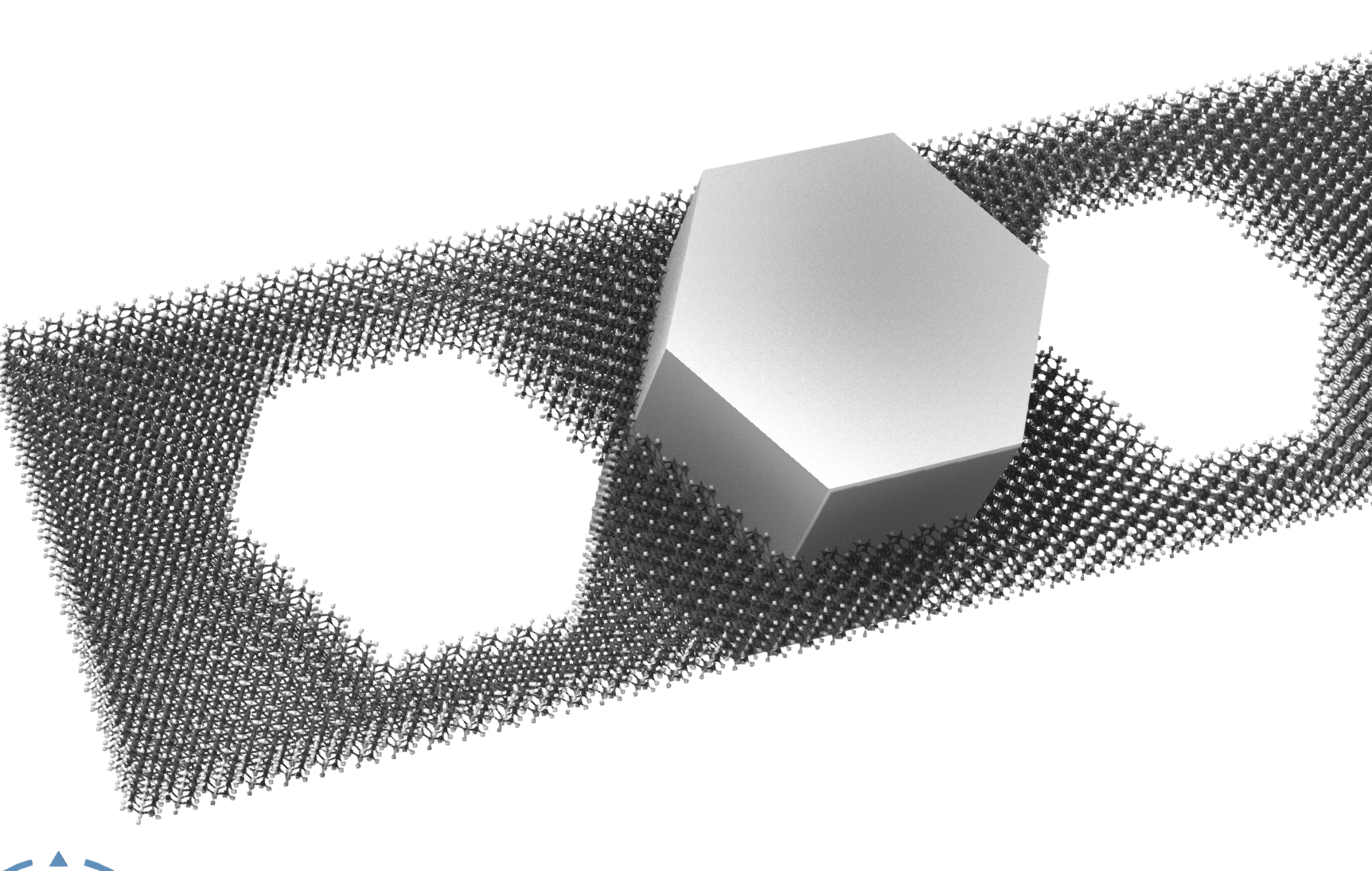

I was wondering if it was possible to use a .stl object to carve crystals in SAMSON. I've been trying out different python snippets provided by the AI assistant which seem like they are just on the verge of doing it, but ultimately are missing some capability I can't quite find a way to add. SAMSON_STL.sam
The idea is it you could position a solid mesh object on your crystal model, do some a test to see if an atom is inside the mesh, and then delete that atom. For simple meshes the test could be if it is inside the bounding box of the mesh.
This could be a cool feature. SAMSON file attached of the system in the image.
Thanks,
Tom
-
I went back and tried to open the file I uploaded here and I get an error that it can't be loaded. Maybe .stls cannot be saved in a .sam?
-
Hi Tom, it might be possible to use some computational geometry libraries that would determine whether a point in inside a triangulated object (it's non trivial for non-convex objects, but they can often be decomposed into convex parts). I'll look into the serialization of the STL objects but I don't immediately see any issues.
-
This one might help: https://github.com/quant-sci/compute-geometry. If you have a convex object like the one in the image, I guess you'd first compute its convex hull thanks to this library, and you'd then use the resulting convex hull to determine whether an atom is inside or not.
-
Thanks for the replies and guidance. I was looking at a couple external libraries to work with stls. I got stuck because I couldn't find anyway to access the stl object through the SAMSON python api then I found Mesh = NodeType.VisualModelMesh? I will work on it again.
-
I think I meant"
from samson import SBNode # in the embedded console it's already imported
Mesh = SBNode.Type.Mesh # or: Mesh = SBNode.Mesh -
The STL importer is actually creating an object of its own class, and cannot be manipulated like a mesh. I'm going to modify that importer so it produces meshes instead. At the moment, you can import obj files instead. However, we also need to expose surfaces in python (meshes are collections of surfaces), which are the classes which expose the underlying vertices and triangles. Without this, you will not be able to do the geometry tests in python (the API is available in C++ though, see https://documentation.samson-connect.net/developers/latest/api/classSBMVisualModelSurface/)
-
Thank you for the tremendous help!
-
I've updated the STL importer. Opening a STL object into SAMSON will now create meshes instead of the previous type of objects (but loading your previous files should work again and will create the previous type of objects, I was able to import your file). The benefit of meshes is that they're the standard way to have triangulated geometry in SAMSON, and you can move them, rotate them, align them, etc. (and we'll expose the underlying surfaces in Python so we can do geometry tests). The update will be available with SAMSON 2026 R1, which should arrive very soon now.
-
That's awesome! I can't wait to try it out. In addition to the crystal carver project, I've done experiments connecting mesh objects and updating them in near real-time from molecular dynamics data. You can see an example here: https://www.youtube.com/watch?v=vMpnWHdG2Ds I actually have a good use case for this. I wonder if anyone else does? Also in the future SAMSON could be used for systems that span from quantum modeling through classical molecular dynamics and into rigid-body-FEA-Multiphysics simulations. I am not aware of any other platform that can do that.
-
Multiscale is definitely worth looking into. Dmitriy exposed surfaces in Python, so here's a script you'll be able to use to carve. It takes the first mesh of the document, and creates two selection groups: one for the atoms inside the mesh, and one for the atoms outside the mesh. It assumes the mesh is watertight to determine whether a point is inside or not. It's not optimized at all but it does the job. (and this script will only work starting with 2026 R1).
import math # ------------------------------------------------------------ # Configuration # ------------------------------------------------------------ MESH_NSL = 'node.type mesh' ATOM_NSL= 'node.type atom' EPS = 1.0e-9 # geometric tolerance # Use a ray direction that is very unlikely to hit many edges exactly RAY_DIR = (1.0, 0.3713906763541037, 0.2182178902359924) # ------------------------------------------------------------ # Small vector utilities # ------------------------------------------------------------ def v_add(a, b): return (a[0] + b[0], a[1] + b[1], a[2] + b[2]) def v_sub(a, b): return (a[0] - b[0], a[1] - b[1], a[2] - b[2]) def v_dot(a, b): return a[0]*b[0] + a[1]*b[1] + a[2]*b[2] def v_cross(a, b): return ( a[1]*b[2] - a[2]*b[1], a[2]*b[0] - a[0]*b[2], a[0]*b[1] - a[1]*b[0], ) def v_scale(a, s): return (a[0]*s, a[1]*s, a[2]*s) def v_norm(a): return math.sqrt(v_dot(a, a)) def v_normalize(a): n = v_norm(a) if n < EPS: return a return (a[0]/n, a[1]/n, a[2]/n) # ------------------------------------------------------------ # Accessors / conversions # ------------------------------------------------------------ def atom_to_point(atom): return (atom.getX().value, atom.getY().value, atom.getZ().value) def position_to_point(position): return (position.x.value, position.y.value, position.z.value) # ------------------------------------------------------------ # Mesh extraction # ------------------------------------------------------------ def extract_triangles_from_mesh(mesh): """ Build a flat triangle list [(v0, v1, v2), ...] from all surfaces of a mesh. Assumes the positions returned by the surfaces are already in the same coordinate frame as the atoms. If your mesh carries a non-identity spatial transform, apply that transform to the vertices before classification. """ triangles = [] transform = mesh.getTransform() for surface in mesh.getSurfaces(): pos = surface.getPositionData() idx = surface.getIndexData() n_pos = int(len(surface.getPositionData())/3) n_tri = int(len(surface.getIndexData())/3) verticesPreTransform = [ SBPosition3(SBQuantity.pm(pos[3*i]), SBQuantity.pm(pos[3*i + 1]), SBQuantity.pm(pos[3*i + 2])) for i in range(n_pos) ] verticesPostTransform = [ transform.transformPoint(verticesPreTransform[i]) for i in range(n_pos) ] vertices = [ position_to_point(verticesPostTransform[i]) for i in range(n_pos) ] for t in range(n_tri): i0 = int(idx[3*t]) i1 = int(idx[3*t + 1]) i2 = int(idx[3*t + 2]) triangles.append((vertices[i0], vertices[i1], vertices[i2])) return triangles # ------------------------------------------------------------ # Point / triangle predicates # ------------------------------------------------------------ def point_on_triangle(p, a, b, c, eps=EPS): """ Check whether p lies on triangle abc within tolerance. """ ab = v_sub(b, a) ac = v_sub(c, a) ap = v_sub(p, a) n = v_cross(ab, ac) nn = v_dot(n, n) if nn < eps: return False # degenerate triangle # Distance from point to triangle plane dist_num = abs(v_dot(ap, n)) if dist_num > eps * math.sqrt(nn): return False # Barycentric test d00 = v_dot(ab, ab) d01 = v_dot(ab, ac) d11 = v_dot(ac, ac) d20 = v_dot(ap, ab) d21 = v_dot(ap, ac) denom = d00 * d11 - d01 * d01 if abs(denom) < eps: return False v = (d11 * d20 - d01 * d21) / denom w = (d00 * d21 - d01 * d20) / denom u = 1.0 - v - w return ( -eps <= u <= 1.0 + eps and -eps <= v <= 1.0 + eps and -eps <= w <= 1.0 + eps ) def ray_intersects_triangle(origin, direction, a, b, c, eps=EPS): """ Moller-Trumbore ray/triangle intersection. Returns distance t along the ray if there is a strict hit, else None. We exclude hits too close to the origin (t <= eps) to avoid self-counting. """ edge1 = v_sub(b, a) edge2 = v_sub(c, a) h = v_cross(direction, edge2) det = v_dot(edge1, h) if abs(det) < eps: return None # parallel or nearly parallel inv_det = 1.0 / det s = v_sub(origin, a) u = inv_det * v_dot(s, h) if u < -eps or u > 1.0 + eps: return None q = v_cross(s, edge1) v = inv_det * v_dot(direction, q) if v < -eps or u + v > 1.0 + eps: return None t = inv_det * v_dot(edge2, q) if t <= eps: return None return t def point_in_watertight_mesh(point, triangles, eps=EPS): """ Parity test: - if the point lies on the surface -> treat as inside - else count ray/triangle intersections """ # Boundary inclusion for a, b, c in triangles: if point_on_triangle(point, a, b, c, eps): return True direction = v_normalize(RAY_DIR) # Collect unique hits; this reduces double-counting when the ray passes # through shared triangle edges or coplanar patches. hits = [] for a, b, c in triangles: t = ray_intersects_triangle(point, direction, a, b, c, eps) if t is not None: hits.append(t) if not hits: return False hits.sort() unique_hits = [] for t in hits: if not unique_hits or abs(t - unique_hits[-1]) > 1.0e-7: unique_hits.append(t) return (len(unique_hits) % 2) == 1 # ------------------------------------------------------------ # Main # ------------------------------------------------------------ mesh_indexer = SAMSON.getNodes(MESH_NSL) if len(mesh_indexer) == 0: raise RuntimeError(f"No mesh found with NSL: {MESH_NSL}") mesh = mesh_indexer[0] print("Mesh:", mesh) triangles = extract_triangles_from_mesh(mesh) print("Number of triangles:", len(triangles)) if len(triangles) == 0: raise RuntimeError("The mesh contains no triangles.") atom_indexer = SAMSON.getNodes(ATOM_NSL) inside_indexer = SBNodeIndexer() outside_indexer = SBNodeIndexer() for atom in atom_indexer: p = atom_to_point(atom) if point_in_watertight_mesh(p, triangles): inside_indexer.addNode(atom) else: outside_indexer.addNode(atom) print("Atoms inside:", inside_indexer.size) print("Atoms outside:", outside_indexer.size) inside_group = SBNodeGroup('inside ' + mesh.name, inside_indexer) outside_group = SBNodeGroup('outside ' + mesh.name, outside_indexer) with SAMSON.holding("Create carving groups"): inside_group.create() SAMSON.getActiveDocument().addChild(inside_group) outside_group.create(); SAMSON.getActiveDocument().addChild(outside_group)
You can then use these groups to delete the atoms you don't want. For example when you delete the atoms in the 'outside' group:
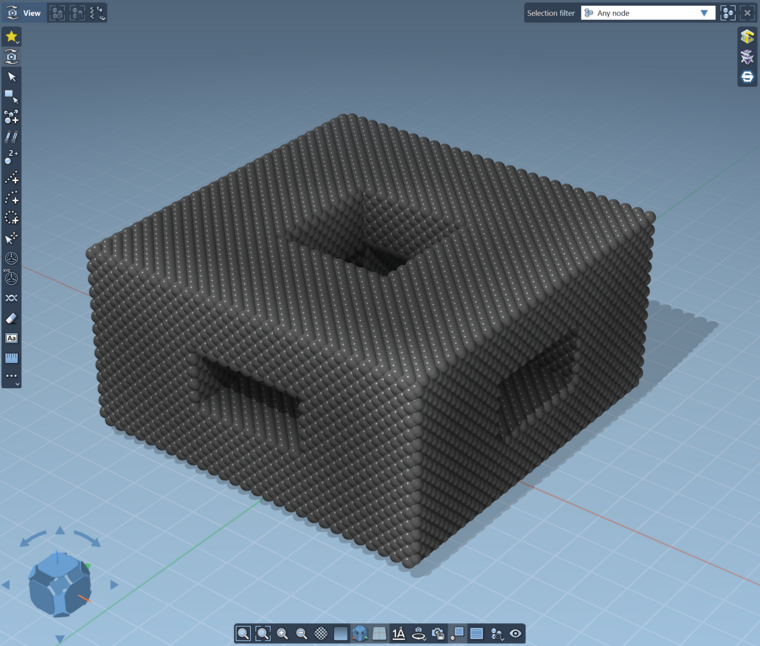
-
This is going to be so cool. It will allow me to make parts so much faster, and parts can be made larger than in NE1. Perhaps Z88 would make a good extension to do exciting stuff with meshes inside SAMSON. https://en.wikipedia.org/wiki/Z88_FEM_software